#include "optimizer/geqo.h"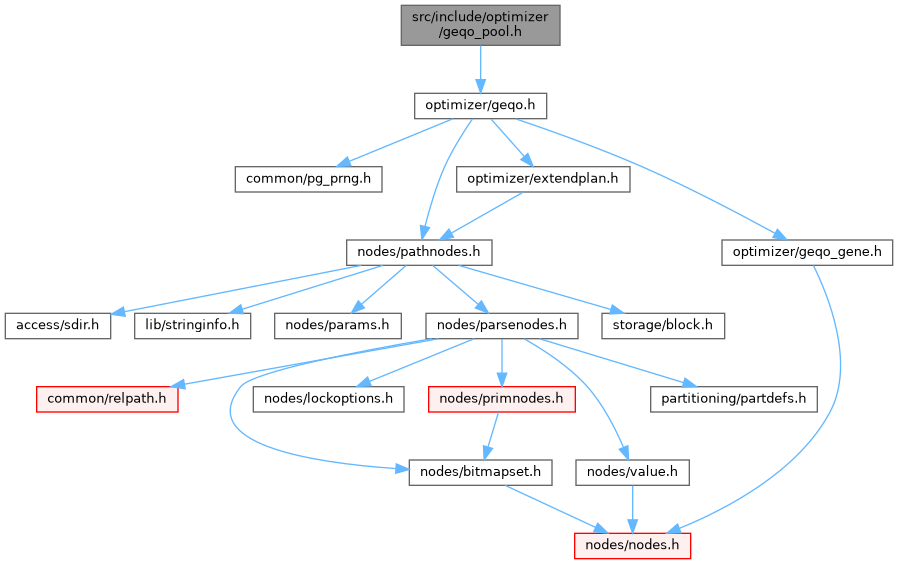
Go to the source code of this file.
Functions | |
| Pool * | alloc_pool (PlannerInfo *root, int pool_size, int string_length) |
| void | free_pool (PlannerInfo *root, Pool *pool) |
| void | random_init_pool (PlannerInfo *root, Pool *pool) |
| Chromosome * | alloc_chromo (PlannerInfo *root, int string_length) |
| void | free_chromo (PlannerInfo *root, Chromosome *chromo) |
| void | spread_chromo (PlannerInfo *root, Chromosome *chromo, Pool *pool) |
| void | sort_pool (PlannerInfo *root, Pool *pool) |
Function Documentation
◆ alloc_chromo()
|
extern |
Definition at line 163 of file geqo_pool.c.
References fb(), palloc_array, and palloc_object.
Referenced by geqo().
◆ alloc_pool()
|
extern |
Definition at line 42 of file geqo_pool.c.
References fb(), i, palloc_array, and palloc_object.
Referenced by geqo().
◆ free_chromo()
|
extern |
◆ free_pool()
|
extern |
Definition at line 69 of file geqo_pool.c.
References Pool::data, fb(), i, pfree(), and Pool::size.
Referenced by geqo().
◆ random_init_pool()
|
extern |
Definition at line 91 of file geqo_pool.c.
References Pool::data, DEBUG1, elog, ERROR, fb(), geqo_eval(), i, init_tour(), root, Pool::size, Pool::string_length, and Chromosome::worth.
Referenced by geqo().
◆ sort_pool()
|
extern |
Definition at line 135 of file geqo_pool.c.
References compare(), Pool::data, qsort, and Pool::size.
Referenced by geqo().
◆ spread_chromo()
|
extern |
Definition at line 190 of file geqo_pool.c.
References Pool::data, fb(), geqo_copy(), i, root, Pool::size, Chromosome::string, Pool::string_length, and Chromosome::worth.
Referenced by geqo().